Haller Lab
The Haller Laboratory is in the Department of Neurosurgery at Washington University School of Medicine and is affiliated with the Departments of Neurology and Genetics.
The laboratory focuses on determining the genetic underpinnings of multiple neurological/neurosurgical disorders including Chiari I malformation, syringomyelia, pediatric hydrocephalus and subarachnoid hemorrhage (SAH). Additionally, we are focused on understanding the functional impact of all potential protein coding genetic variation in neurological disease genes to better predict which patients are at risk for developing these disorders.

Current Research
The overall goal of the Haller laboratory is to identify and characterize genetic variation that leads to various neurosurgical and/or neurological disorders in humans. By collecting and sequencing large cohorts of patients with the same disorder (CM1, SM, SAH, Hydrocephalus), we identify genes and genetic variants that contribute to risk of developing these disorders. We then characterize the genes and genetic variants we find associated with these disorders using cell-culture models and by disrupting disease-associated genes in zebrafish.
Genetics of Chiari I malformation
We have assembled a cohort of >1000 CM1 patients and relatives and are performing exome sequencing to identify genetic variation that we can link to CM1 risk and progression. We leverage the vast infrastructure of the Park-Reeves Syringomyelia Research Consortium to connect with CM1 patients and their families to build the largest cohort of CM1 patients with genetic information in the world. Additionally, we have access to >5000 unrelated control exome sequenced individuals through long-time collaboration at Washington University which we use to compare to CM1 patients. We are leaders in rare variant association methods and are working to determine the role of rare genetic variation in CM1 pathogenesis.
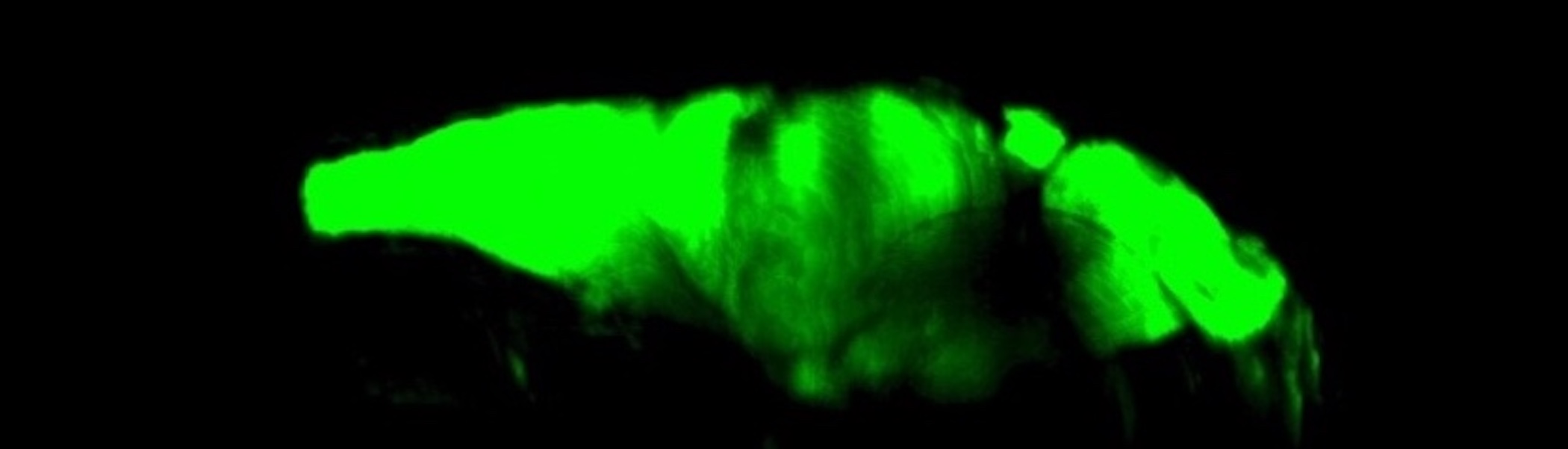
Zebrafish models of Neurological Disorders
We use zebrafish as a model system to study the development of various neurological disorders including Chiari I malformation, syringomyelia and hydrocephalus. By “knocking-out” disease-associated genes in zebrafish, we are able to see what role these genes play in the development of analogous traits in the zebrafish.
High-throughput Functional Studies
We are developing high-throughput methods for dissecting the functional impact of genetic variation in their genes or non-coding regions of interest. The production of libraries of DNA molecules, each different from the reference by one and only one position currently requires the purchase of many thousands of synthesized oligonucleotides either to be directly cloned into a vector, used as the donor DNA with CRISPR-Cas9 or used as primers in multiplex mutagenesis protocol. We have developed a method of massively parallel single nucleotide mutagenesis in which such libraries can be made in a single day for<$30/RXN. This method was published at Nature Methods and we currently hold a patent. We are currently using this method to functionally assess all possible single nucleotide changes in several Mendelian disease genes to predict which variants are capable of causing disease.
Lab Team
- Gabriel Haller, PhD, Principal Investigator
- ebbie Moeller, Lab Technician
- Jackson Wilborn, Lab Technician
- Tim Kuensting, Clinical Research Coordinator
- Geng Wang, Graduate Student
- Amin Munshi, Undergraduate Student
- Brianna Vo, Undergraduate Student
- Jessica Shapiro, Undergraduate Student